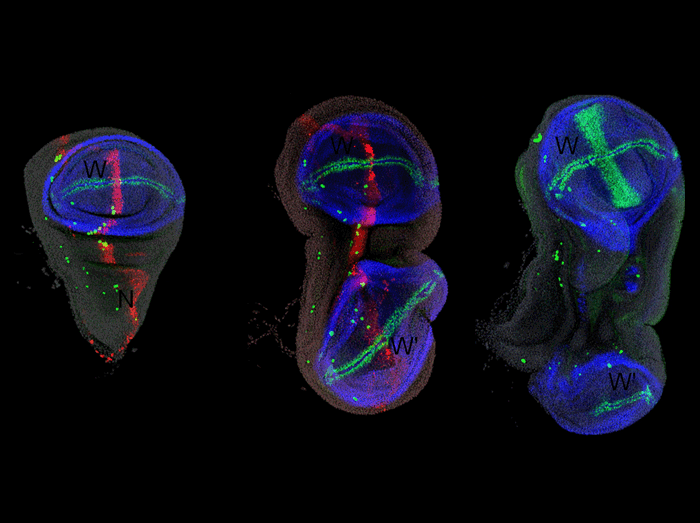2008 Drosophila Image Award
Winner
Global Analysis of mRNA Localization
While mRNA localization is known to play important roles in the establishment of cellular asymmetries, it is generally thought to be a rare phenomenon limited to specialized transcripts. To assess the overall variety and prevalence of mRNA localization events on a genomic level, we conducted a high-resolution FISH analysis of approximately 25% of mRNAs encoded in the Drosophila genome. We found that the majority of sampled mRNAs (71%) are subcellularly localized and that transcripts with similar localization patterns are often functionally related. Some of the striking and diverse patterns observed in the early embryo are illustrated in this journal cover image, with mRNAs depicted in blue and nuclei in red. This study suggests that mRNA localization is a widespread biological mechanism for organizing localized cellular machineries. An accompanying database provides a powerful resource for predicting gene function and for further characterizing mRNA localization pathways.
Eric Lécuyer, Hideki Yoshida, Neela Parthasarathy, Christina Alm, Tomas Babak, Tanja Cerovina, Timothy R. Hughes, Pavel Tomancak, and Henry M. Krause. (2007) Global Analysis or mRNA Localization Reveals a Prominent Role in Organizing Cellular Architecture and Function. Cell 131:174-187.
Runner-up
A microRNA that can activate Wingless signaling
(Left) ptc-Gal4, UAS-DsRed wing disc, stained for DsRed (red), Nubbin (blue) to mark the wing pouch, and Senseless (green) to mark the wing margin and sensory cells. (Center) ptc-Gal4; UAS-DsRed-mir-315 wing disc exhibits duplication of the wing pouch (W) at the expense of the notum (N). (Right) dpp-Gal4, UAS-wg wing disc also exhibits a notum-to-wing transformation.
Silver, S.*, J. W. Hagen*, K. Okamura, N. Perrimon and E. C. Lai (2007). Functional screening identifies miR-315 as a potent activator of Wingless signaling. Proc. Natl. Acad. Sci. USA, 104: 18151-18156. (*, these authors contributed equally).
Runner-Up
Mapping Pheromone and Fruit Odor Representations
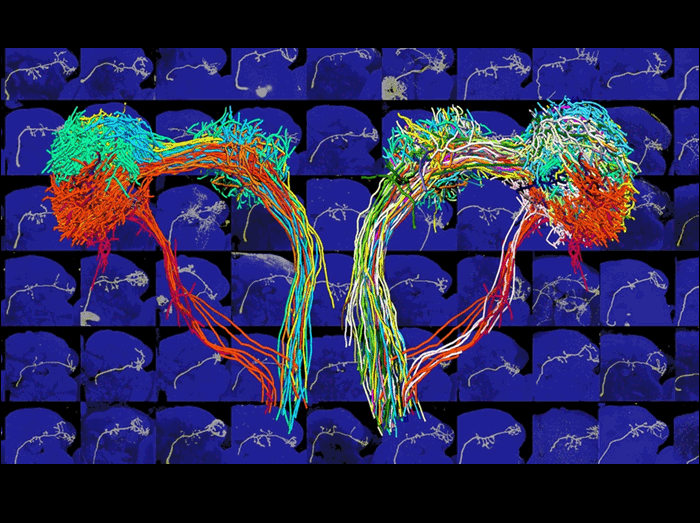
How are external stimuli, such as odors, represented inside the brain? In Drosophila, projection neurons (PNs) carry odor information from the antennal lobes to higher brain centers. In this image, single PNs from hundreds of different brains (some of which are shown in the background) have been assembled onto a single template brain by a process of warping registration, which can map brains of slightly different shapes onto one another.
On the right, 229 PN axons belonging to 33 glomerular classes are labeled by a class-specific color. Each PN class synapses with specific olfactory receptor neurons and sends stereotyped projections to two higher olfactory centers, the mushroom body calyx and lateral horn (the most lateral structure).
The left side shows a subset of PN axons, emphasizing that putative pheromone-representing PNs (red color shades) project their axons to an area of the lateral horn brain region distinct from those that process fruit odors (green color shades).
The lateral horn region of the brain is therefore compartmentalized to reflect biologically relevant olfactory inputs, suggesting how different odor stimuli may be coupled to distinct behavioral outputs.
Jefferis GS, Potter CJ, Chan AM, Marin EC, Rohlfing T, Maurer CR Jr, Luo L.
Comprehensive maps of Drosophila higher olfactory centers: spatially segregated fruit and pheromone representation. Cell128,187-203 (2007)
Finalist
Morphogen Precision
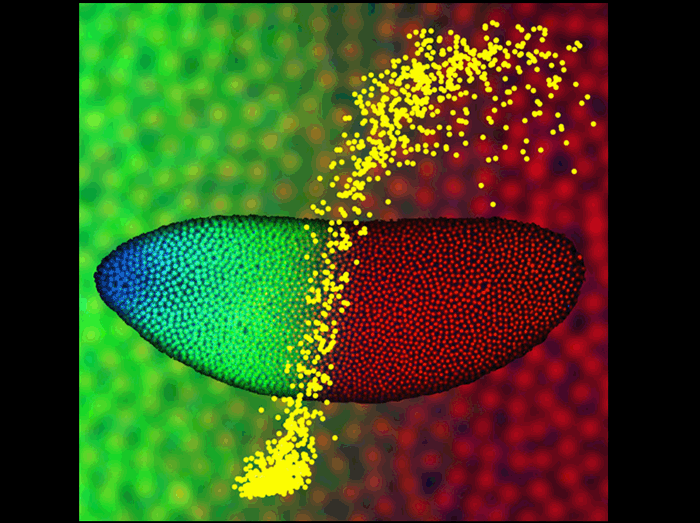
The image shows a Drosophila embryo 2 hr after fertilization, with nuclei at the surface fluorescently labeled for Bicoid protein (blue), Hunchback protein (green), and DNA (red). Using two-photon microscopy these embryos were imaged to quantitatively characterize the dynamics and precision of how morphogen molecules communicate positional information to individual nuclei. In this example, the shallow Bicoid gradient generates a sharp Hunchback boundary (enlarged in the background), partitioning the embryo in half. This input/output relationship is quantitatively represented in the foreground (yellow), where each dot specifies the Bicoid concentration (horizontal axis) and Hunchback concentration (vertical axis) measured in a single nucleus. The results indicate that the precision with which the embryo interprets and locates this boundary is very high, approaching limits set by simple physical principles.
Thomas Gregor, David W. Tank, Eric F. Wieschaus, and William Bialek,
Probing the Limits to Positional Information, Cell 130, 153-164 (2007).
Finalist
Wing pattern evolution in Hawaiian Drosophila
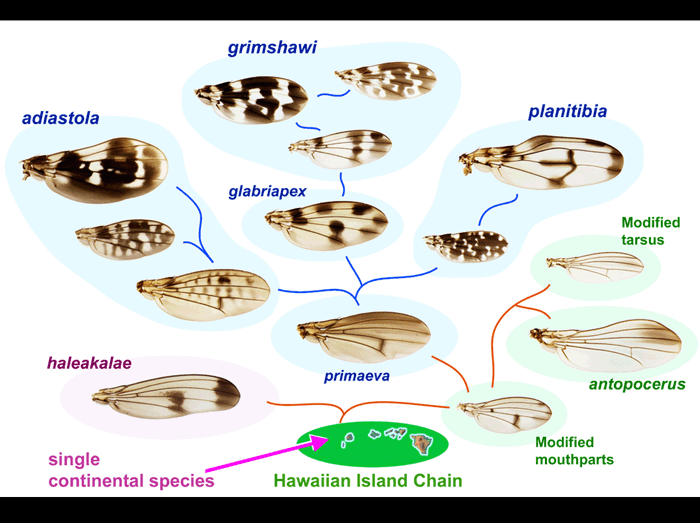
There are 500-1000 remarkably diverse species of Drosophila endemic to Hawaii. In this paper, wings of 73 of these species were documented and compared. Here, a sample of 14 species is used to illustrate the proposed phylogeny of the Hawaiian radiation.
All the species appear to be descended from a single introduction of Drosophila to the Hawaiian Island chain. These flies diverged into several groups characterized by distinct secondary sexual characters, such as male-specific modified mouthparts or legs, and other features. (Names of the main species groups are shown.)
The Picture Wing species (blue backgrounds) include multiple independent lineages that exhibit increasing complexity of wing pigment pattern, as shown.
Edwards, K.A., L.T. Doescher, K.Y. Kaneshiro, and D. Yamamoto. 2007. A database of wing diversity in the Hawaiian Drosophila.PLoS ONE 2(5): e487, 1-12.
Finalist
Color-coded mitotic spindles of Drosophila S2 cells
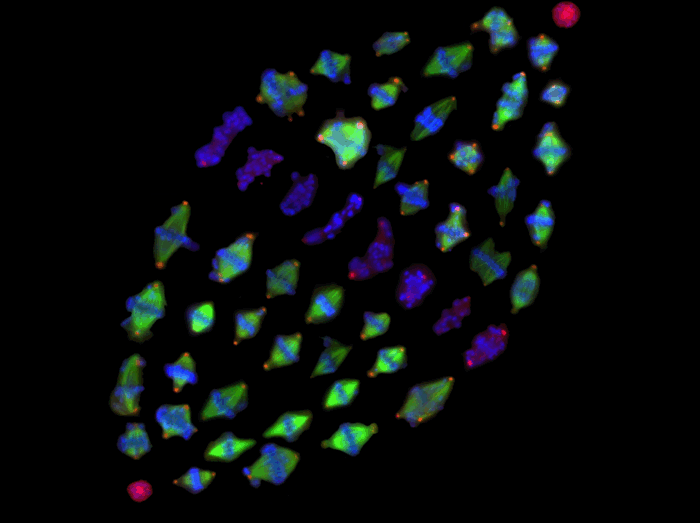
The formation of a metaphase spindle, a bipolar microtubule array with centrally aligned chromosomes, is a prerequisite for the faithful segregation of a cell's genetic material. Using a full-genome RNAi screen of Drosophila S2 cells, we identified ~200 genes that contribute to spindle assembly. This figure is composed of high-throughput microscopy images of abnormal-shaped metaphase spindles that are derived from 51 different RNAi treatments. The phenotypes include multipolar, anastral or longer/shorter spindles and misaligned chromosomes.
Gohta Goshima, Roy Wollman, Sarah S. Goodwin, Nan Zhang, Jonathan M. Scholey, Ronald D. Vale, Nico Stuurman.
Genes required for mitotic spindle assembly in Drosophila S2 cells
Science 316: 417-21 (2007)
Finalist
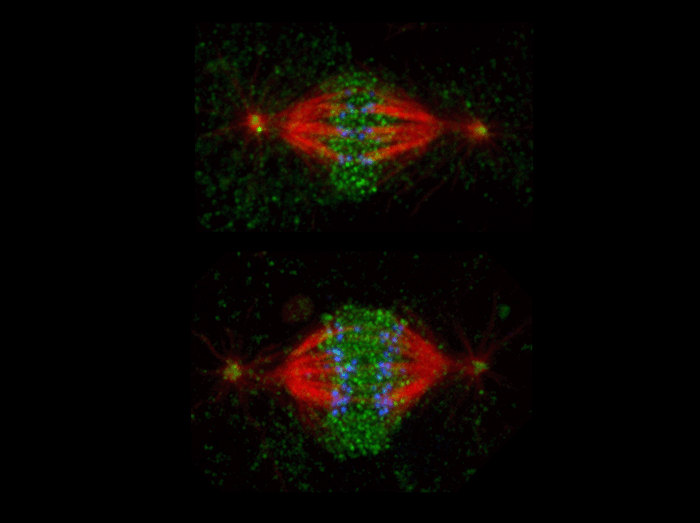
The microtubule severing protein, katanin, localizes to chromosomes and regulates anaphase A
The movement of chromosomes on the mitotic spindle requires the depolymerization of microtubule ends. The figure shows the mitotic localization of the microtubule severing protein katanin (green) relative to spindle microtubules (red) and kinetochores/chromosomes (blue). Katanin targets to chromosomes during both metaphase (top) and anaphase (bottom) and is responsible for inducing the depolymerization of attached microtubule plus-ends. We have proposed that katanin does so specifically by removing stabilizing plus-end “caps”.
Dong Zhang, Gregory C. Rogers, Daniel W. Buster, and David J. Sharp. Three microtubule severing proteins contribute to the “Pacman-Flux” machinery that moves chromosomes. The Journal of Cell Biology, 177, 231-242 (2007).
Finalist
Tissue replacement in cell competition depends on cell engulfment
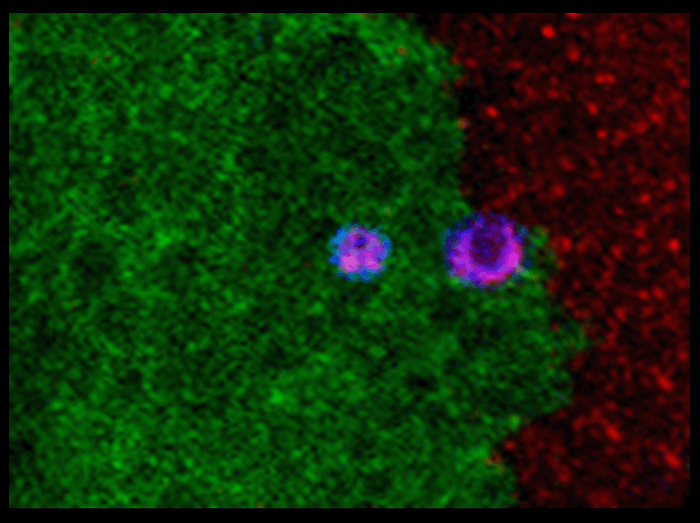
When two genotypes of cell are present in a chimeric individual, cell competition can lead the favored genotype to replace the other during organ growth.
This image of a wing disc shows that wild type epithelial cells (green) generated by mitotic recombination from Minute cells (red) are replacing the later cells by cell competition. This process depends on and could be induced by a phagocytic-dependent cell-killing process, in which wild type cells actively kill Minute cells. Consequently, the dead Minute cells (magenta) are engulfed and replaced by the killing wild type cells.
It indicates that tissue replacement, when involving cell competition, may be achieved by an approach including engulfment manipulation, without altering cells’ intrinsic growth rates.
Wei Li and Nicholas E. Baker. 2007. Engulfment is required for cell competition. Cell, 129(6):1215-1225
Finalist
Egfr/Ras signaling affects epithelial cell morphology in the Drosophila wing
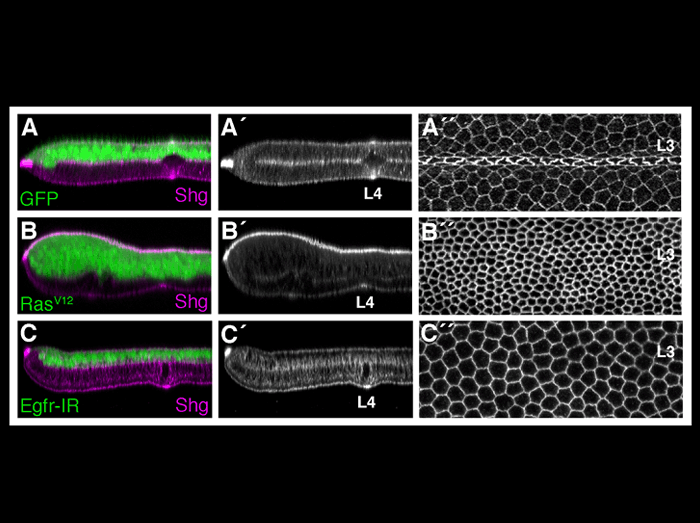
During the pupal stage of development, vein and intervein cells of the Drosophila wing become morphologically distinct. While the importance of Ras/Egfr signaling in vein cell fate determination has been extensively documented, the mechanisms linking Ras activity to vein differentiation have been unclear. This figure demonstrates that the Egfr/Ras signaling pathway affects the apical/basal distribution of DE-cadherin/Shotgun to affect epithelial cell morphology. Compared to wildtype wings (A) ectopic activation of Egfr/Ras signaling (B) promotes apical constriction (as in vein cells), whereas loss of Egfr activity (C) promotes cell flattening (as in intervein cells). Optical cross sections through vein L4 (to the left) or the dorsal wing surface of vein L3 (to the right) are shown. All wings are stained with an antibody specific for DE-cadherin/Shotgun.
O’Keefe, D.D., Prober, D.A., Moyle, P.S., Rickoll, W.L., Edgar, B.A. (2007). Egfr/Ras signaling regulates DE-cadherin/Shotgun localization to control vein morphogenesis in the Drosophilawing. Dev Biol 311, 25-39.
Finalist
Mother centrosome is inherited by the stem cell
Drosophila male germ line stem cells always divide asymmetrically in the context of signaling in the niche. Such asymmetric stem cell division is ensured by the stereotyped movement of centrosomes that prepares perpendicularly oriented spindle toward the niche. We have found that difference(s) between mother and daughter centrosomes underlie such an asymmetric stem cell division: The mother centrosome always stays close to the niche, while the daughter migrates away from the niche, being inherited by a differentiating daughter, gonialblast. This is the first demonstration that mother and daughter centrosomes behave differently in the context of tissue architecture/development.
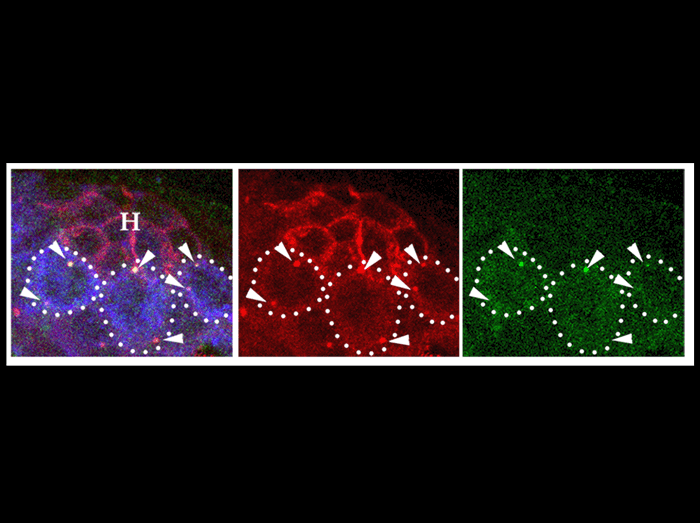
The image shows a testis tip from newly eclosed male with GSCs containing GFP-labeled mother centrosomes proximal to the hub-GSC interface. To label mother centrosomes, a centriolar marker, GFP-PACT, was expressed only during early embryogenesis by using NGT40 driver. Red ; -tubulin (centrosomes, arrowheads) and Fas III (hub, H). Green ; GFP-PACT. Blue; Vasa (germ cells). Bar 10µm.
Yamashita YM, Mahowald AP, Perlin JR, Fuller MT. Asymmetric inheritance of mother versus daughter centrosome in stem cell division.Science. 2007 Jan 26;315(5811):518-21.
Finalist
mirtrons define a novel class of microRNA precursor
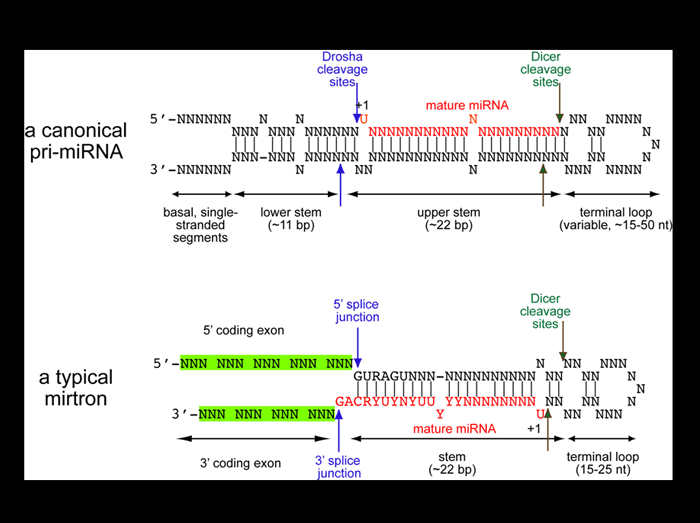
Analysis of cloned Drosophila small RNAs revealed a novel family of regulatory RNAs derived from short intronic hairpins termed “mirtrons”. Unlike canonical miRNAs, whose extended stem directs their cleavage by the nuclear RNAse III enzyme Drosha (blue arrows), mirtrons are bounded by coding exons (green) and are defined directly by the splicing machinery (blue arrows). Both types of hairpins are then cleaved in the cytoplasm by Dicer to yield 21-22 nt inhibitory RNAs. The recognition of Drosophila mirtrons led to the identification of an analogous class of mammalian mirtron genes.
K. Okamura, J. W. Hagen, H. Duan, D. Tyler and E. C. Lai (2007). The mirtron pathway generates microRNA-like regulatory RNAs in Drosophila. Cell 130: 89-100.